Romain Lannes
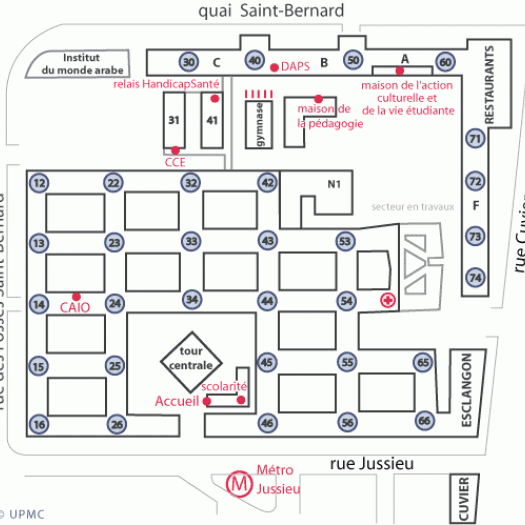

14 heures - Tour 46/56 - 2eme étage
J’ai le plaisir de vous inviter à la soutenance de ma thèse qui s’intitule « Recherche de séquences environnementales inconnues d’intérêt médical/biologique par l’utilisation de grands réseaux de similarité de séquences ». La thèse sera suivi d’une collation ou vous êtes cordialement invités.
Composition du jury :
Sakina-Dorothée Ayata (Examinatrice),
Eric Bapteste (directeur),
Chris Bowler (Examinateur),
Claudine Landès (Rapportrice),
Catherine Larose (Rapportrice),
Philippe Lopez (directeur).
–
The objective of this thesis was to identify as yet unknown microorganisms present in various environments and to characterize some of their metabolisms. This unidentified diversity, both taxonomic and functional, is commonly referred to as microbial dark matter. I have used and developed new network methods, including sequence similarity networks, to exploit very large sequence datasets from metagenomic projects. In particular, my work has highlighted the ecological role of ultra-small micro-organisms in some autotrophic metabolic pathways in the oceans. It also shows that CPR and DPANN, recently discovered ultra-small bacteria and archaea, participate in the dynamics of microbial communities through quorum sensing systems similar to those of better-characterized organisms. An application of sequence similarity networks to meta-barcoding data also revealed a previously unknown diversity of Holozoans, which could allow us to better understand the transition to multicellularity of Metazoans. Finally, I have developed a method and software for searching for remote homologs of proteins of interest in very large datasets, such as those from metagenomics. This method, now validated, should make it possible to search for sequences belonging to still unknown and very divergent organisms, in the hope of discovering new deep branching phyla, or even new domains of life.